An R6Class of AIPW for estimating the average causal effects with users' inputs of exposure, outcome, covariates and related libraries for estimating the efficient influence function.
Value
AIPW object
Details
An AIPW object is constructed by new() with users' inputs of data and causal structures, then it fit() the data using the
libraries in Q.SL.library and g.SL.library with k_split cross-fitting, and provides results via the summary() method.
After using fit() and/or summary() methods, propensity scores and inverse probability weights by exposure status can be
examined with plot.p_score() and plot.ip_weights(), respectively.
If outcome is missing, analysis assumes missing at random (MAR) by estimating propensity scores of I(A=a, observed=1) with all covariates W.
(W.Q and W.g are disabled.) Missing exposure is not supported.
See examples for illustration.
Constructor
AIPW$new(Y = NULL, A = NULL, W = NULL, W.Q = NULL, W.g = NULL, Q.SL.library = NULL, g.SL.library = NULL, k_split = 10, verbose = TRUE, save.sl.fit = FALSE)
Constructor Arguments
| Argument | Type | Details |
Y | Integer | A vector of outcome (binary (0, 1) or continuous) |
A | Integer | A vector of binary exposure (0 or 1) |
W | Data | Covariates for both exposure and outcome models. |
W.Q | Data | Covariates for the outcome model (Q). |
W.g | Data | Covariates for the exposure model (g). |
Q.SL.library | SL.library | Algorithms used for the outcome model (Q). |
g.SL.library | SL.library | Algorithms used for the exposure model (g). |
k_split | Integer | Number of folds for splitting (Default = 10). |
verbose | Logical | Whether to print the result (Default = TRUE) |
save.sl.fit | Logical | Whether to save Q.fit and g.fit (Default = FALSE) |
Constructor Argument Details
W,W.Q&W.gIt can be a vector, matrix or data.frame. If and only if
W == NULL,Wwould be replaced byW.QandW.g.Q.SL.library&g.SL.libraryMachine learning algorithms from SuperLearner libraries or
sl3learner object (Lrnr_base)k_splitIt ranges from 1 to number of observation-1. If k_split=1, no cross-fitting; if k_split>=2, cross-fitting is used (e.g.,
k_split=10, use 9/10 of the data to estimate and the remaining 1/10 leftover to predict). NOTE: it's recommended to use cross-fitting.save.sl.fitThis option allows users to save the fitted sl object (libs$Q.fit & libs$g.fit) for debug use. Warning: Saving the SuperLearner fitted object may cause a substantive storage/memory use.
Public Methods
| Methods | Details | Link |
fit() | Fit the data to the AIPW object | fit.AIPW |
stratified_fit() | Fit the data to the AIPW object stratified by A | stratified_fit.AIPW |
summary() | Summary of the average treatment effects from AIPW | summary.AIPW_base |
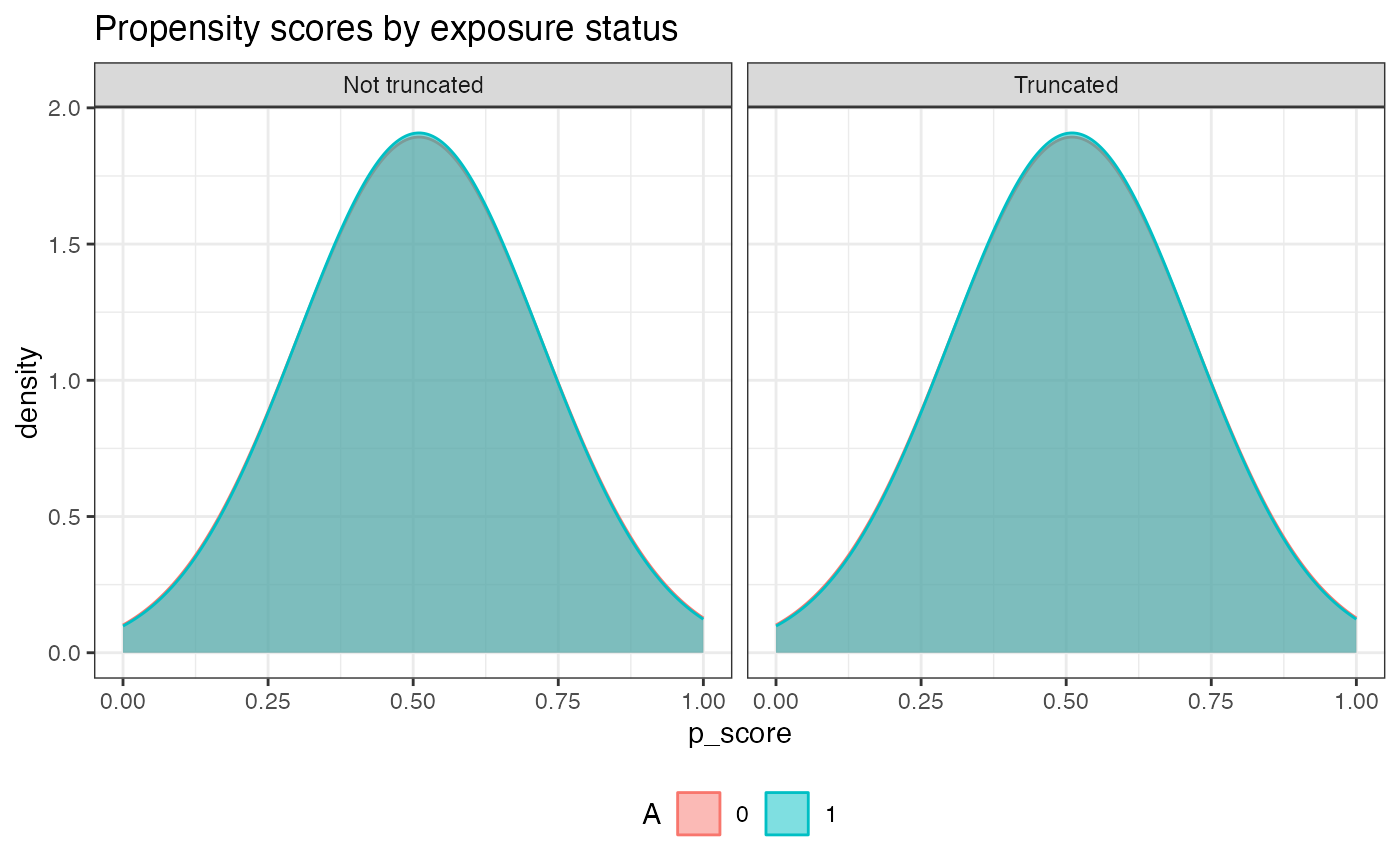
plot.p_score() | Plot the propensity scores by exposure status | plot.p_score |
plot.ip_weights() | Plot the inverse probability weights using truncated propensity scores | plot.ip_weights |
Public Variables
| Variable | Generated by | Return |
n | Constructor | Number of observations |
stratified_fitted | stratified_fit() | Fit the outcome model stratified by exposure status |
obs_est | fit() & summary() | Components calculating average causal effects |
estimates | summary() | A list of Risk difference, risk ratio, odds ratio |
result | summary() | A matrix contains RD, ATT, ATC, RR and OR with their SE and 95%CI |
g.plot | plot.p_score() | A density plot of propensity scores by exposure status |
ip_weights.plot | plot.ip_weights() | A box plot of inverse probability weights |
libs | fit() | SuperLearner or sl3 libraries and their fitted objects |
sl.fit | Constructor | A wrapper function for fitting SuperLearner or sl3 |
sl.predict | Constructor | A wrapper function using sl.fit to predict |
Public Variable Details
stratified_fitAn indicator for whether the outcome model is fitted stratified by exposure status in the
fit()method. Only when usingstratified_fit()to turn onstratified_fit = TRUE,summaryoutputs average treatment effects among the treated and the controls.obs_estAfter using
fit()andsummary()methods, this list contains the propensity scores (p_score), counterfactual predictions (mu,mu1&mu0) and efficient influence functions (aipw_eif1&aipw_eif0) for later average treatment effect calculations.g.plotThis plot is generated by
ggplot2::geom_densityip_weights.plotThis plot uses truncated propensity scores stratified by exposure status (
ggplot2::geom_boxplot)
References
Zhong Y, Kennedy EH, Bodnar LM, Naimi AI (2021). AIPW: An R Package for Augmented Inverse Probability Weighted Estimation of Average Causal Effects. American Journal of Epidemiology.
Robins JM, Rotnitzky A (1995). Semiparametric efficiency in multivariate regression models with missing data. Journal of the American Statistical Association.
Chernozhukov V, Chetverikov V, Demirer M, et al (2018). Double/debiased machine learning for treatment and structural parameters. The Econometrics Journal.
Kennedy EH, Sjolander A, Small DS (2015). Semiparametric causal inference in matched cohort studies. Biometrika.
Examples
library(SuperLearner)
#> Loading required package: nnls
#> Loading required package: gam
#> Loading required package: splines
#> Loading required package: foreach
#> Loaded gam 1.22-2
#> Super Learner
#> Version: 2.0-28.1
#> Package created on 2021-05-04
library(ggplot2)
#create an object
aipw_sl <- AIPW$new(Y=rbinom(100,1,0.5), A=rbinom(100,1,0.5),
W.Q=rbinom(100,1,0.5), W.g=rbinom(100,1,0.5),
Q.SL.library="SL.mean",g.SL.library="SL.mean",
k_split=1,verbose=FALSE)
#fit the object
aipw_sl$fit()
# or use `aipw_sl$stratified_fit()` to estimate ATE and ATT/ATC
#calculate the results
aipw_sl$summary(g.bound = 0.025)
#check the propensity scores by exposure status after truncation
aipw_sl$plot.p_score()